Sequence Alignment and Clustering
This example compares evolutionarily preserved (homologous) gene sequences between different organisms, using sequence alignment and clustering.
The EME1 gene has homologs across many species. The Wolfram Knowledgebase provides EME1 sequence data for humans, chimpanzees, mice and rats to both Wolfram|Alpha and the Wolfram Language.
For example, here is the the EME1 gene sequence for mice.
Now interpret EME1 gene entities for each of the selected organisms.
From here, retrieve the gene sequences for each of the selected genes.
Looking at the beginnings of the sequence alignments, it appears that the EME1 gene sequences for humans and chimpanzees are much more similar than those for rats and mice. Human and mice EME1 sequences appear even less similar.
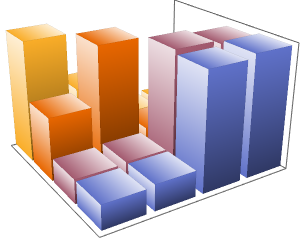
With sequence clustering, you will be able to see how similar the entire sequences actually are. First, calculate all similarity ratios between all of the sequences.
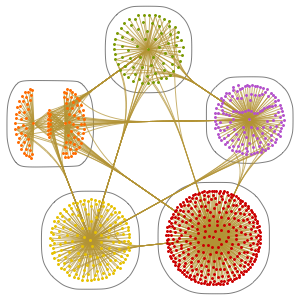
You can see strong similarities between the human and chimpanzee chromosomes and medium similarities between the rat and mouse chromosomes.
By taking the dendrogram, you can see the relative tightness of the clusters produced from this similarity.
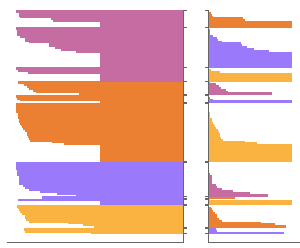
Looking across the entire alignment, you can see that human EME1 sequences are nearly perfectly aligned to chimpanzee EME1 sequences, while the EME1 alignment between mice and humans is close to random.
With music, we can hear the difference between stronger versus weaker alignment, signifying in music where the sequences are aligned versus when they differ.
Listen for yourself to see how the chimpanzee/human alignment for EME1 sounds.
Compare that to the sound of the EME1 alignment between rats and mice.