Drug Dose Selection
Therapeutic monoclonal antibodies (mAbs) are becoming an increasingly important class of drugs in the treatment of many severe diseases. Due to their very specific nature and long half-lives, however, selecting safe, first-in-human starting doses for new mAbs can be tricky. In this example, we will see how a mathematical modeling approach, using Wolfram SystemModeler and Mathematica, can be used to aid such a dose selection process.
The Model
For this new drug, let us assume that a target occupancy (TO) of 10% is the desired goal, meaning that the administered dose should result in only 10% of the target molecules being bound to the mAb. What drug dose does this goal correspond to?
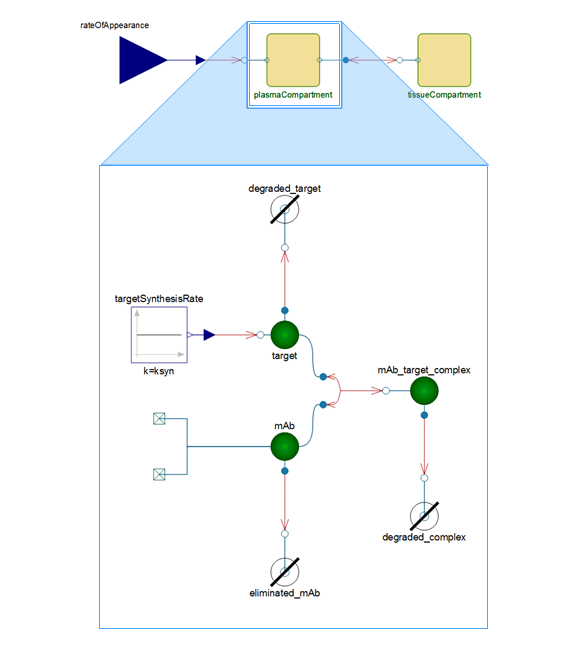
A target-mediated drug disposition model is used to describe the pharmacokinetic and pharmacodynamic behavior of the drug. The tissue compartment represents nonspecific tissue binding or distribution, while the plasma compartment displays the path of the mAb from injection to binding and finally to direct elimination or degradation. The plasma compartment also includes the synthesis and degradation of the target molecule, which both affect the TO.
Finding a Safe Drug Dose
By using Mathematica and Wolfram SystemModeler Link, it is possible to identify the maximum TO for each simulation and thereby create a plot of how the maximum TO changes with the chosen drug dosage.
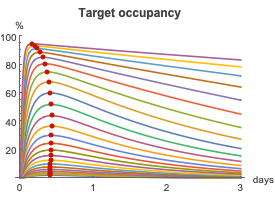
By calculating the TO for a multitude of different doses, we can cover a wide range of maximum TOs, from no target molecules being occupied up to 100% occupation. The red dots indicate the maximum TO for each dose and are used to generate the plot to the right.
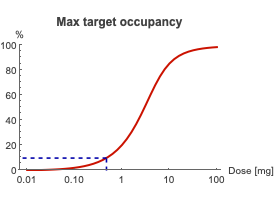
Dose response curve indicating the dose that corresponds to a maximum TO of 10%.
Wolfram System Modeler
Try
Buy
System Modeler is available in English
and Japanese
on Windows, macOS & Linux »
Questions? Comments? Contact a Wolfram expert »