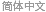Find a Splice Site in a DNA Sequence
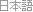
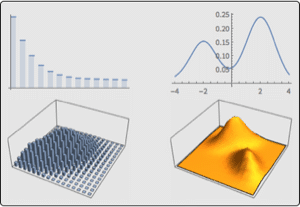
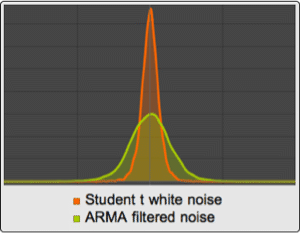
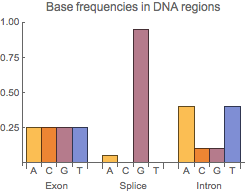
A DNA sequence consists of base letters A, C, G, and T. Suppose there is a sequence that begins in an exon, contains a splice site, and ends in an intron. If the exons have a uniform base composition, the introns are deficient in C and G, and the splice site consensus nucleotide is G with probability 0.95, the frequency distributions are as follows.
| In[1]:= |  X |
 |
| In[2]:= | X |
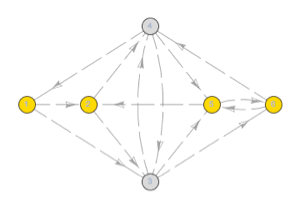
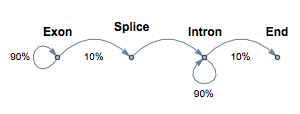
The state machine has states for exon (1), splice (2), intron (3), and end (4), with the following transition probabilities between states.
 |
| In[3]:= | X |
The emissions are nucleotides A (1), C (2), G (3), T (4), or end (5).
| In[4]:= | X |
| In[5]:= | X |
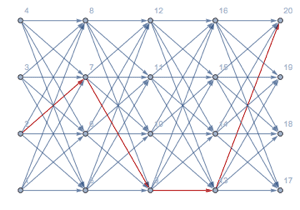
Find the most probable nucleotide subsequence (exon, splice, intron, or end).
| In[6]:= | X |
| Out[6]= |
Find the joint probability of the preceding nucleotide sequence and the DNA sequence.
| In[7]:= | X |
| Out[7]= |